FlyMAD - The Fly Mind Altering Device
Today I'm proud to announce the availability of all source code, and the advanced online publication of our paper
Bath DE*, Stowers JR*, Hörmann D, Poehlmann A, Dickson BJ, Straw AD (* equal contribution) (2014) FlyMAD: Rapid thermogenetic control of neuronal activity in freely-walking Drosophila. Nature Methods. doi 10.1038/nmeth.2973
FlyMAD (Fly Mind Altering Device) is a system for targeting freely walking flies (Drosophila) with lasers. This allows rapid thermo- and opto- genetic manipulation of the fly nervous system in order to study neuronal function.
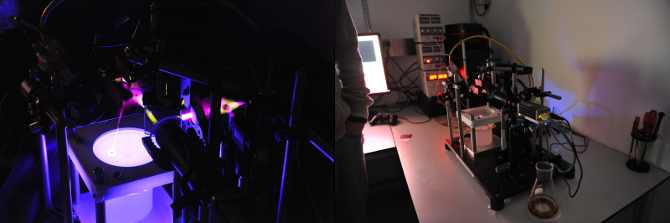

The scientific aspects of the publication are better summarised on nature.com, here, on our laboratory website, or in the video at the bottom of this post.
Briefly however; if one wishes to link function to specific neurons one could conceive of two broad approaches. First, observe the firing of the neurons in real time using fluorescence or other microscopy techniques. Second, use genetic techniques to engineer organisms with light or temperature sensitive proteins bound to specific neuronal classes such that by the application of heat or light, activity in those neurons can be modulated.
Our system takes the second approach; our innovation being that by using real time computer vision and control techniques we are able to track freely walking Drosophila and apply precise (sub 0.2mm) opto- or thermogenetic stimulation to study the role of specific neurons in a diverse array of behaviours.
This blog post will cover a few of the technical and architectural decisions I made in the creation of the system. Perhaps it is easiest to start with a screenshot and schematic of the system in operation
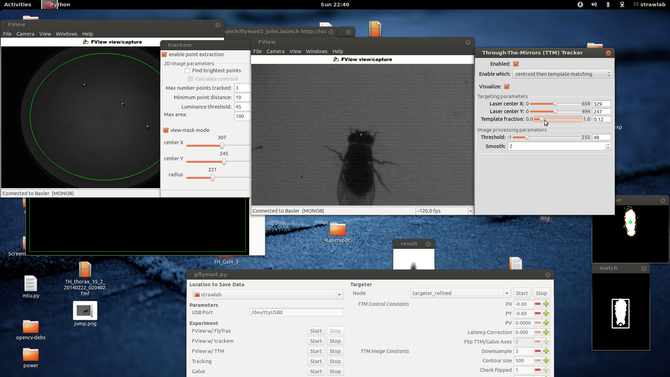
Here one can see two windows showing images from the two tracking cameras, associated image processing configuration parameters (and their results, at 120fps). In the center at the bottom is visible the ROS based experimental control UI. Schematically, the two cameras and lasers are arranged like the following
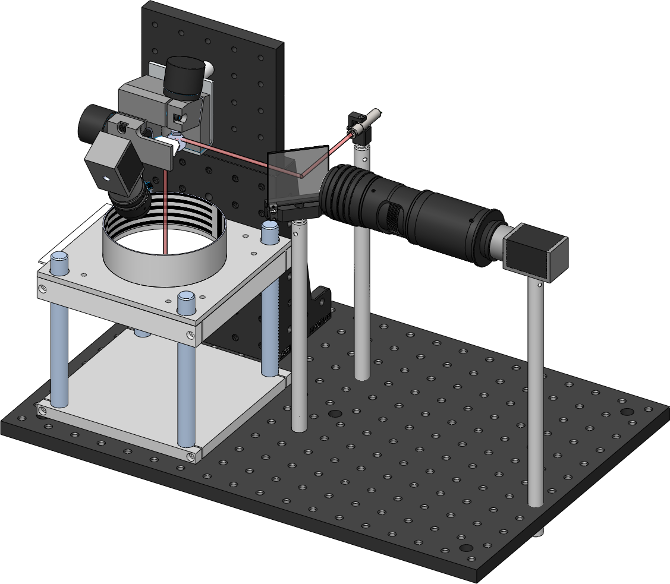
In this image you can also see the thorlabs 2D galvanometers (top left), and the dichroic mirror which allows aligning the camera and laser on the same optical axis.
By pointing the laser at flies freely walking in the arena below, one can subsequently deliver heat or light to specific body regions.
General Architecture
The system consists of hardware and software elements. A small microcontroller and digital to analogue converter generate analog control signals to point the 2D galvanometers and to control laser power. The device communicates with the host PC over a serial link. There are two cameras in the system; a wide camera for fly position tracking, and a second high magnification camera for targeting specific regions of the fly. This second camera is aligned with the laser beam, and its view can be pointed anywhere in the arena by the galvanometers.
The software is conceptually three parts; image processing code, tracking and targeting code, and experimental logic. All software elements communicate using robot operating system (ROS) interprocess communication layer. The great majority of code is written in python.
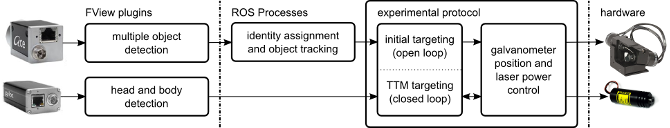
Robot Operating System (ROS)
ROS is a framework traditionally used for building complex robotic systems. In particular it has a relatively good performance and simple, strongly typed, inter-process-communication framework and serialization format.
Through its (pure) python interface one can build a complex system of multiple processes who communicate (primarily) by publishing and subscribing to message "topics". An example of the ROS processes running during a FlyMAD experiment can be seen below.
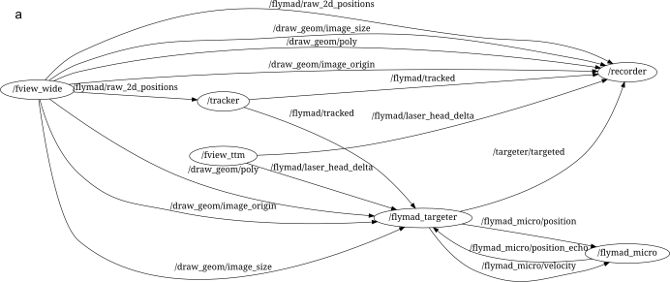
The lines connecting the nodes represent the flow of information across the network, and all messages can be simultaneously recorded (see /recorder) for analysis later. Furthermore, the isolation of the individual processes improves robustness and defers some of the responsibility for realtime performance from myself / Python, to the Kernel and to my overall architecture.
For more details on ROS and on why I believe it is a good tool for creating reliable reproducible science, see my previous post, my Scipy2013 video and presentation
Image Processing
There are two image processing tasks in the system. Both are implemented as FView plugins and communicate with the rest of the system using ROS.
Firstly, the position of the fly (flies) in the arena, as seen by the wide camera, must be determined. Here, a simple threshold approach is used to find candidate points and image moments around those points are used to find the center and slope of the fly body. A lookup table is used to point the galvanometers in an open-loop fashion approximately at the fly.
With the fly now located in the field of view of the high magnification camera a second real time control loop is initiated. Here, the fly body or head is detected, and a closed loop PID controller finely adjusts the galvanometer position to achieve maximum targeting accuracy. The accuracy of this through the mirror (TTM) system asymptotically approaches 200μm and at 50 msec from onset the accuracy of head detection is 400 ± 200 μm. From onset of TTM mode, considering other latencies in the system (gigabit ethernet, 5 ms, USB delay, 4 ms, galvanometer response time, 7 ms, image processing 8ms, and image acquisition time, 5-13 ms) total 32 ms, this shows the real time targeting stabilises after 2-3 frames and comfortably operates at better than 120 frames per second.
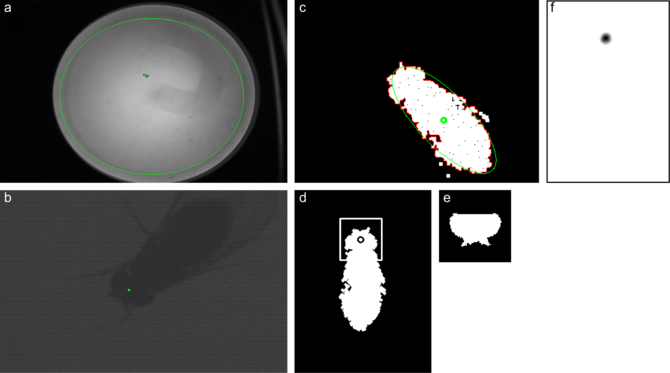
To reliably track freely walking flies, the head and body step image processing operations must take less than 8ms. Somewhat frustratingly, a traditional template matching strategy worked best. On the binarized, filtered image, the largest contour is detected (c, red). Using an ellipse fit to the contour points (c,green), the contour is rotated into an upright orientation (d). A template of the fly (e) is compared with the fly in both orientations and the best match is taken.
I mention the template strategy as being disappointing only because I spent considerable time evaluating newer, shinier, feature based approaches and could not achieve the closed loop performance I needed. While the newer descriptors, BRISK, FREAK, ORB were faster than the previous class, nether (in total) were significantly more reliable considering changes in illumination than SURF - which could not meet the <8ms deadline reliably. I also spent considerable time testing edge based (binary) descriptors such as edgelets, or edge based (gradient) approaches such as dominant orientation templates or gradient response maps. The most promising of this class was local shape context descriptors, but I also could not get the runtime below 8ms. Furthermore, one advantage of the contour based template matching strategy I implemented, was that graceful degradation was possible - should a template match not be found (which occurred in <1% of frames), an estimate of the centre of mass of the fly was still present, which still allowed degraded targeting performance. No such graceful fallback was possible using feature correspondence based strategies.
There are two implementations of the template match operation - GPU and CPU based. The CPU matcher uses the python OpenCV bindings (and numpy in places), the GPU matcher uses cython to wrap a small c++ library that does the same thing using OpenCV 2.4 Cuda GPU support (which is not otherwise accessible from python). Intelligently, the python OpenCV bindings use numpy arrays to store image data, so passing data from Python to native code is trivial and efficient.
I also gave a presentation comparing different strategies of interfacing python with native code. The provided source code includes examples using python/ctypes/cython/numpy and permutations thereof.
The GPU code-path is only necessary / beneficial for very large templates and higher resolution cameras (as used by our collaborator) and in general the CPU implementation is used.
Experimental Control GUI
To make FlyMAD easier to manage and use for biologists I wrote a small GUI using Gtk (PyGObject), and my ROS utility GUI library rosgobject.
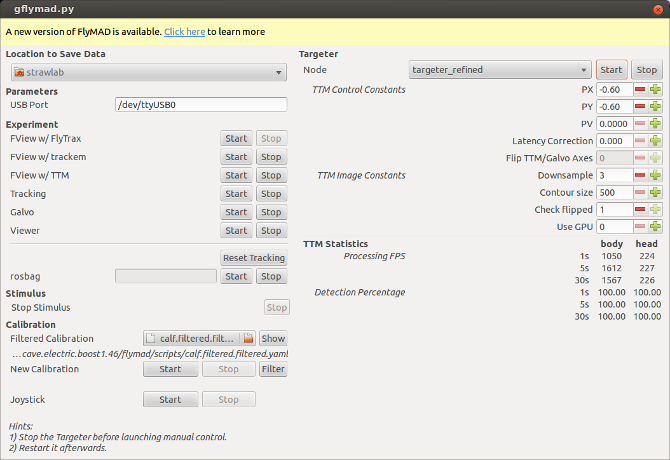
On the left you can see buttons for launching individual ROS nodes. On the right are widgets for adjusting the image processing and control parameters (these widgets display and set ROS parameters). At the bottom are realtime statistics showing the TTM image processing performance (as published to ROS topics).
Like good ROS practice, once reliable values are found for all adjustable parameters they can be recorded in a roslaunch file allowing the whole system to be started with known configuration from a single command.
Manual Scoring of Videos
For certain experiments (such as courtship) videos recorded during the experiment must be watched and behaviours must be manually annotated. To my surprise, no tools exist to make this relatively common behavioural neuroscience task any easier (and easier matters; it is not uncommon to score 10s to 100s of hours of videos).
During every experiment, RAW uncompressed videos from both cameras are written to disk (uncompressed videos are chosen for performance reasons, because SSDs are cheap, and because each frame can be precisely timestamped). Additionally, rosbag files record the complete state of the experiment at every instant in time (as described by all messages passing between ROS nodes). After each experiment finishes, the uncompressed videos from each camera are composited together, along with metadata such as the frame timestamp, and a h264 encoded mp4 video is created for scoring.
After completing a full day of experiments one can then score / annotate videos in bulk. The scorer is written in Python, uses Gtk+ and PyGObject for the UI, and vlc.py for decoding the video (I chose vlc due to the lack of working gstreamer PyGObject support on Ubuntu 12.04)
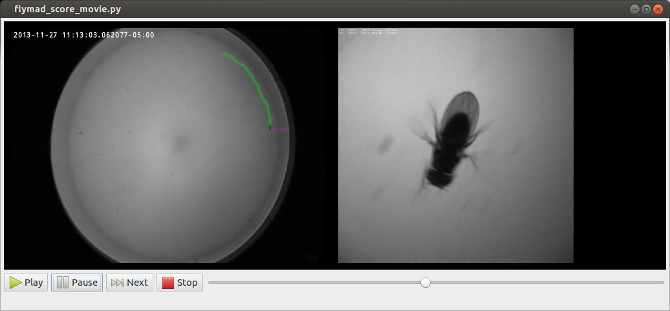
In addition to allowing play, pause and single frame scrubbing through the video, pressing any of qw,as,zx,cv pairs of keys indicates that a a behaviour has started or finished. At this instant the current video frame is extracted from the video, and optical-character-recognition is performed on the top left region of the frame in order to extract the timestamp. When the video is finished, a pandas dataframe is created which contains all original experimental rosbag data, and the manually annotated behaviour against on a common timebase.
Distributing complex experimental software
The system was not only run by myself, but by collaborators, and we hope in future, by others too. To make this possible we generate a single file self installing executable using makeself, and we only officially support one distribution - Ubuntu 12.04 LTS and x86_64.
The makeself installer performs the following steps
- Adds our Debian repository to the system
- Adds the official ROS Debian repository to the system
- Adds our custom ROS stacks (FlyMAD from tarball and rosgobject from git) to the ROS environment
- Calls rosmake flymad to install all system dependencies and build and non-binary ROS packages.
- Creates a FlyMAD desktop file to start the software easily
We also include a version check utility in the FlyMAD GUI which notifies the user when a newer version of the software is available.
The Results
Using FlyMAD and the architecture I have described above we created a novel system to perform temporally and spatially precise opto and thermogenetic activation of freely moving drosophila. To validate the system we showed distinct timing relationships for two neuronal cell types previously linked to courtship song, and demonstrated compatibility of the system to visual behaviour experiments.
Practically we were able to develop and simultaneously operate this complex real-time assay in two countries. The system was conceived and built in approximately one year using Python. FlyMAD utilises many best-in-class libraries and frameworks in order to meet the demanding real time requirements (OpenCV, numpy, ROS).
We are proud to make the entire system available to the Drosophila community under an open source license, and we look forward to its adoption by our peers.
For those still reading, I encourage you to view the supplementary video below, where its operation can be seen.
Comments, suggestions or corrections can be emailed to me or left on Google Plus